The Extreme-Scale Scientific Software Stack (E4S): A New Resource for Computational and Data Science Research
By: Rob Farber, contributing writer
The extreme-scale scientific software stack (E4S) provides a fundamentally new resource for computational and data science research communities. As an integrated, curated and tested portfolio of open-source, reusable scientific software components, E4S provides building blocks for the scientific community. Application developers can build directly on top of E4S, gaining access to the latest software capabilities and dramatically reducing the time and effort to manage dependencies. Computing facilities and vendors can use E4S as a resource for their software stack, reducing the time and cost to support important components. Developers of reusable scientific software can contribute their work to E4S, making it available to a broad user base. Scientists get access to tested and verified software components, algorithms and heuristics. E4S runs on laptops and the latest supercomputers, and everything in between. E4S is a new member of the HPC ecosystem, making real the potential for fundamental advances in productivity and sustainability in service of the advance of science and engineering.
Readers interested in learning more and developers who wish to join E4S can visit the E4S Project GitHub community or download the latest ECP container from http://E4S.io (instructions are provided at the end of this article). The E4S Build Cache currently has 22,292 packages available for a variety of Linux distributions.
At a meta level, E4S saves developers time and reduces users’ agony. E4S is a unified portal for buildable, scalable, robust, tested, and verified software with associated forums along with containers that can benefit developers, scientists, and vendors. More than that, E4S complements existing capabilities, such as GitHub, while also acting as a central collaborative tool to stimulate discussion, continuous improvement, and the dissemination of timely information on everything from best practices to new algorithms, documentation, bug fixes, and new features.
“E4S is open and inclusive,” says Dr. Mike Heroux, Director of Software Technology (ST) for the ECP. “Projects can join and become first-class citizens simply by showing that the software can be built and satisfy the core community policies.” In addition to his role with the ECP, Dr. Heroux is a senior scientist at Sandia National Laboratories and a scientist in residence at St. John’s University. “We are still building community,” he adds, “and we invite any interested members of the HPC and scientific communities to join us as partners to provide excellent software both effectively and efficiently.”
E4S is open and inclusive. Projects can join and become first-class citizens simply by showing that the software can be built and satisfy the core community policies. We are still building community and we invite any interested members of the scientific and HPC communities to join us as partners to provide excellent software both effectively and efficiently.
—Dr. Mike Heroux, Director of Software Technology for the ECP
How to read this article
E4S is extensive. For this reason, this article begins by highlighting two examples of how the E4S initiative already contributes value to many Python and OpenMP programmers. These examples are provided to show the immediate opportunity presented by E4S.
After the use cases, a summary view of E4S is provided along with references to more detailed information about the ECP capabilities. These references are intended to be used as an index to direct readers to their particular areas of interest. A simple example that readers can download to start the ECP Singularity container is provided at the end of this article to show how easy it is to try the software.
Already delivering immediate benefits to developers and researchers
Dr. Lois Curfman McInnes, Senior Computational Scientist at Argonne National Laboratory and Deputy Director of ST for the ECP, points out that E4S is already delivering immediate and actionable benefits. “E4S is already benefitting the HPC and scientific communities. Some current HPC efforts rely on up to 20 of the currently 90 E4S projects,” she says.
E4S is already benefitting the HPC and scientific communities. Some current HPC efforts rely on up to 20 of the currently 90 E4S projects.
—Dr. Lois Curfman McInnes, Senior Computational Scientist at Argonne National Laboratory and Deputy Director of ST for the ECP
Use case: Distributed GPU-accelerated Python using NumPy with Legate
Python is a highly popular programming language. Used as the implementation language in Gordon Bell finalist projects,[i] Python is now considered a first-class citizen in HPC. NumPy is a popular Python library used for performing array-based numerical computations. Such operations are common in scientific codes as well as finance, business, computer aided design, and more.
Legate, a member of E4S, is a programming system that transparently accelerates and distributes NumPy programs to machines of any scale and capability that are typically used by changing one module import statement.[ii]
Legate works by translating NumPy programs to the Legion programming model and then leverages the scalability of the Legion run time system to distribute data and computations across an arbitrary-sized machine. Compared with similar programs written in the distributed Dask array library in Python, Legate has demonstrated that it can achieve speedups of up to 10× on 1,280 CPUs and 100× on 256 GPUs.[iii]
Use case: Optimized OpenMP on GPUs
E4S projects are being designed to benefit all programmers, regardless of whether programmers are coding for HPC or industry. For example, OpenMP is arguably the most prevalent multithreading programming model in use today. By using a programming construct called a pragma, programmers can annotate their C, C++, or Fortran program to use all the cores and vector units on a modern multicore CPU or offload the computational work to an accelerator such as a GPU. This can result in a substantial performance increase.[iv]
SOLLVE[v],[vi] is an ECP project that is tasked with delivering high-performance OpenMP codes for improved scalability and message passing interface (MPI) interoperability on DOE exascale supercomputers. SOLLVE(Scaling OpenMP with LLVM for Exascale Performance and Portability) is currently heavily focused on offload features for devices such as NVIDA and AMD GPUs. The SOLLVE project, which is part of E4S, has already delivered a 32.1% performance boost over the current LLVM11 release on an ECP application that was in the 2020 ECP OpenMP hackathon—specifically, the GridMini lattice QCD benchmark.[vii] Even such an early performance boost can reduce the time-to-solution for an HPC project by one-third.
Even more exciting, Vivek Kale, a computer scientist at Brookhaven National Laboratory, says, “SOLLVE is unique in its view of the GPU as a collection of streaming multiprocessor units (SMs) rather than a monolithic device. This fits naturally with the OpenMP threading model.” To put this in perspective, a SOLLVE task parallel loop on a GPU with 128 SMs can run up to 128× small problems concurrently on each SM. Subject to operating system (OS) restrictions, this means that an OpenMP task parallel loop can—in theory, at least—run 128× faster by using task parallelism compared with an implementation based on a monolithic single instruction, multiple data execution model that would run each task sequentially on a GPU.[viii]
A validated and tested software repository led by established authorities
E4S is a curated and regular release of ECP ST products built by using Spack.
Spack is an open-source package manager geared toward simplified, reproducible builds of the otherwise complicated dependency chains common in HPC software, and it provides the ability to leverage existing compilers and run time system libraries for native software installations. E4S also provides validation verification test suites to ensure that software packages build correctly and can run a validation test suite. Several Linux OS and machine combinations are supported, including x86, ARM, and PPC processor platforms as well as GPU-accelerated systems (currently AMD and NVIDIA).
The scope of E4S can be seen in the 90 projects already targeted for integration and release. As shown in Figure 1, this includes libraries from xSDK (the Extreme-scale Scientific Software Development Kit), such as hypre, MAGMA, MFEM, PETSc/TAO, SUNDIALS, SuperLU, and Trilinos. Additional libraries and packages are being seriously considered for addition to the suite.
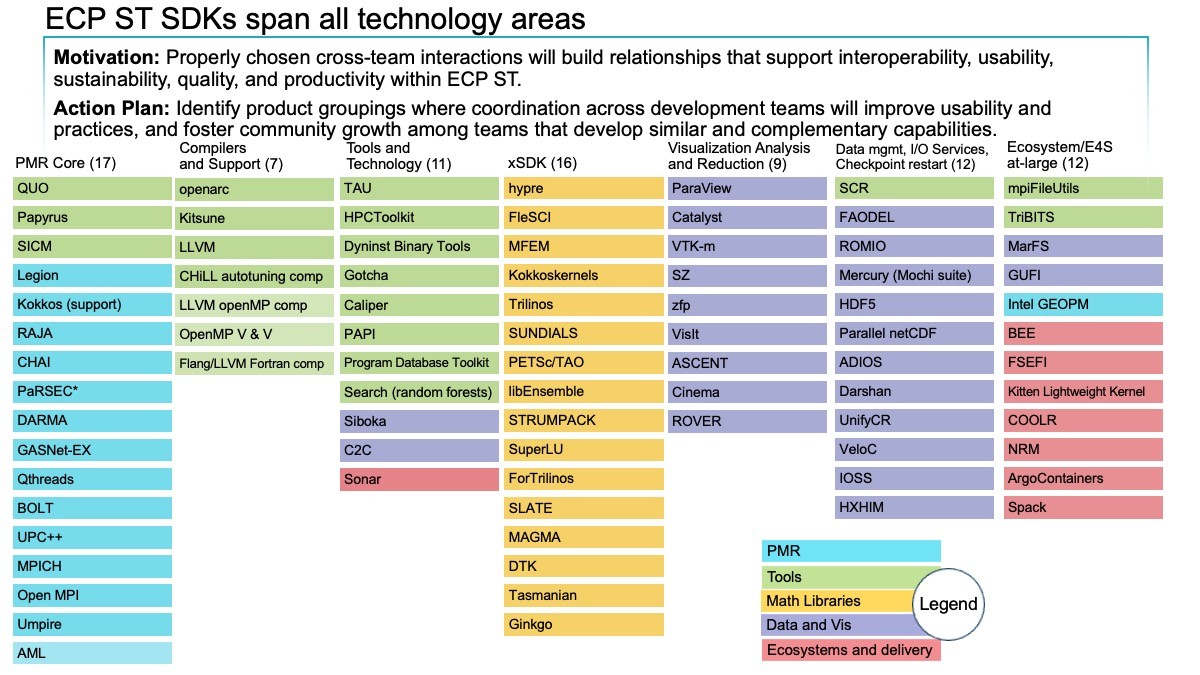
Figure 1: As of February 2020, the ECP ST products are organized into six software development kits (SDKs) (first six columns). The rightmost column lists products that are not part of an SDK but are part of the Ecosystem group that will also be delivered as part of E4S. The colors denoted in the key map all the ST products to the ST technical area of which they are a part. For example, the xSDK consists of products that are in the Math Libraries Technical area, plus TuckerMPI, which is in the Ecosystem and Delivery technical area. (Source: ECP Software Technology Capability Assessment Report)
The E4S product portfolio is led by accomplished and recognized team leaders in their respective fields, as shown in Figure 2.

Figure 2: The ECP ST leadership team.
Finding more in one’s areas of interest
The ECP Software Technology Capability Assessment Report contains much information about the current state of E4S. The current incarnation is very long and is too much to absorb at one time.
Figure 1 shows that the ECP ST currently has six level 3 (L3) technical areas. An L3 technical area is an identified area that is funded to provide guidance and oversight for a set of sub-projects. Each technical area funds specific level 4 technology projects within their respective focus area. For example, Programming Models and Runtimes has a group working on an Exascale MPI/MPICH, Mathematical Libraries has a group working on PETSc, and so forth. To learn more, readers can download the ECP Software Technology Capability Assessment Report, then go to Section 4 in the table of contents and click on the specific technology project that is of interest.
Figure 3 provides an example TOC entry for an ECP ST technical area.

Figure 3: Example TOC entry for an ECP ST technical area.
Figure 4 illustrates where the different projects fit in the software stack relative to the hardware.
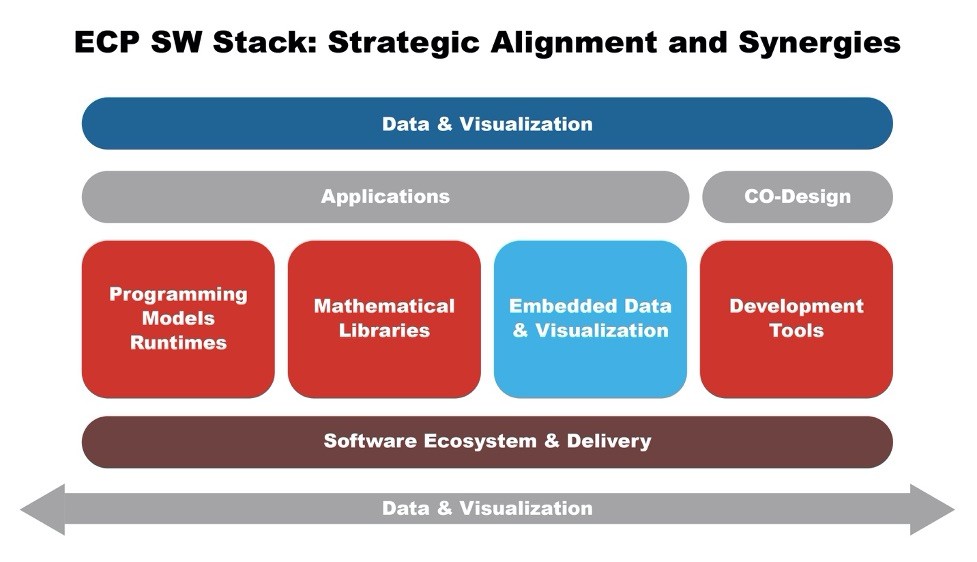
Figure 4: Illustration of where the ECP SW stack products reside in the software stack relative to the hardware.
Along with software and tools, E4S provides a collaborative environment
Although software and tools are a key focus, E4S also provides a collaborative environment in which researchers can work together to address the myriad of software issues associated with running at scale. Success relies on more than just a foundation based on high-quality, tested software. Success also relies on enabling people to use the software. This is why Dr. Heroux describes the documentation portal as “a beautiful thing.”
The documentation portal is a beautiful thing.
—Dr. Mike Heroux, Director of ST for the ECP
The E4S vision covers the HPC and scientific software life cycle
Dr. McInnes explains how the DOE exascale effort created the vision for E4S. “Never before has there been such a broad and deep focus on quality HPC software. Along with supporting the forthcoming US exascale supercomputers and accelerated-node computing in general, our ambition is to become a community effort spanning a variety of national and international organizations that also includes vendors,” she says. Conversations about common challenges and goals are under way with the National Science Foundation; NASA; various government labs and academic institutions; commercial organizations, such as HPE; and the Hartree Institute in the United Kingdom, to name a few. Technology transfer to vendors is a desirable outcome that can benefit consumers and scientific and HPC communities by helping hardware vendors, software developers, systems integrators, cloud providers, enterprise datacenters, and hyperscalers.
Never before has there been such a broad and deep focus on quality HPC software.
—Dr. Lois Curfman McInnes, Senior Computational Scientist at Argonne National Laboratory and Deputy Director of ST for the ECP
As a collaborative effort, E4S covers all aspects of the scientific software life cycle, including building from source, testing, documentation, deployment (including facilities specific deployments), and workflow and data management.
User forums are also an integral part of E4S. According to Dr. Heroux, “The opportunity to incorporate software best practices and interoperability to enable collaboration is unprecedented. Never before has there been this opportunity and at this scale.”
Rethinking HPC is necessary
The breadth of E4S is necessary to meet the unique demands of scientific computing, now and in the future. Architecture changes at all levels of computing challenge traditional thinking and force innovation. The impact affects accelerated-node computing in general from laptops to desktops to high-end and leadership systems.
Even on current systems, scientists are challenged because their data are too big to move without consuming inordinate amounts of time and effort. This seemingly simple, generic runtime challenge will only worsen as scientific workflows use increasingly powerful computers in academic institutions and datacenters around the world. Such basic challenges demonstrate why E4S is needed: to help HPC and scientific communities rethink their tools and workflows, as well as system design, data storage, and data warehousing.
Rethinking tools and workflows benefit everyone. For example, the current seismic shift in visualization workflows that resulted from increased data size occurred with the advent of in situ data reduction, analysis, and in-transit scientific visualization. These workflows are becoming popular because they minimize and can even completely eliminate data movement by rendering the data on the same nodes that perform the computation. Eliminating data movement makes real-time photorealistic rendering possible, even when rendering extremely large datasets and putatively exascale-sized datasets. [i], [ii], [iii], [iv], [v]
The E4S website offers much more than what is discussed in this article and is rapidly evolving to meet community needs. As the website states, “E4S exists to reduce barriers for testing, distributing, and using open-source software capabilities needed for advancing computational science and engineering.”
Try an E4S container today
Readers can try an E4S container today by downloading the correct container for their system. E4S v1.2 contains 67 E4S products in a full featured Singularity container for x86_64 and ppc64le platforms. Click the “Download” button on the E4S.io website or go to https://e4s-project.github.io/download.html. Then, perform the following steps.
1. Select the right container for your system (i.e., either Docker or Singularity and the correct machine/OS configuration).
2. If using the latest version of Singularity, then type:
singularity exec [download container name].simg /bin/bash
Designed to be used in production
The current container is around 13 GB. Such a large container size can be frowned upon by systems management teams.
Sameer Shende, Director of the Performance Research Lab of the University of Oregon’s Neuroinformatics Center, says, “We are working on an E4S-cl container launch tool that will transparently replace the MPI in the application compiled with the software stack in the container with the system MPI to get near-native inter-node communication performance. This tool uses the singularity —sandbox option to expand (untar) the image so the run time loader just reads the set of libraries that are required by the application and does not need to load/read the full image during the launch of a job. This can potentially speedup the execution of container-based workflow considerably. We also have base and full featured images on DockerHub.”
Applications can customize their image by using the base images that feature: (1) support for GPUs and (2) MPI, compilers, and bare-bones tools, such as vi and make, to create a compact image. The second approach is detailed in the use of E4S for Pantheon and WDMapp, and references are provided in the next section.
Anecdotal performance reports reflect the performance potential. David Rogers, Team Lead for Data Science at Scale Team at Los Alamos National Laboratory, observes that, “For the current workflows, we get approximately 10×speedup in Spack build-from-source vs. Spack install-from-cache (~1 min vs. ~12 min). We are timing only the Spack build/install of Ascent and its dependent packages and at the moment are not including the build-time of the application (Nyx), which is not currently cached.”
For more information
The following are a targeted set of resources that provide more information.
- For a more detailed description, see the whitepaper E4S: Extreme-scale Scientific Software Stack.
- The E4S GitHub has several whitepapers.
- WorkshopResources provides recordings, posters, reports, and more.
- The highlights and slides for Pantheon are available from the University of Oregon Dropbox:
- The E4S section for WMDapp is described at https://wdmapp.readthedocs.io/en/latest/machines/rhea.html;search for “E4S.”
Rob Farber is a global technology consultant and author with an extensive background in HPC and machine learning technology development that he applies at national labs and commercial organizations. Rob can be reached at [email protected].
[i] https://www.osti.gov/servlets/purl/1567661
[ii] https://www.osti.gov/biblio/1558501
[iii] https://woiv.gitlab.io/woiv17/ISC_WOIV_Keynote1_1_Jeffers.pdf
[i] https://www.olcf.ornl.gov/2016/08/23/streamlining-accelerated-computing-for-industry/
[ii] https://legion.stanford.edu/pdfs/legate-preprint.pdf
[iii] ibid
[iv] For example, a highly parallel code could realize a speedup of several-cores per computational node over a sequential code. Modern processors can have 48 cores per socket. A typical HPC system has two processors per computational node. On such a system, an OpenMP code can theoretically realize a 2 × 48 or 96× speedup. Efficient use of the vector units can deliver an 8× or greater additional speedup on floating-point and integer-intensive parallel loops.
[v] https://www.exascaleproject.org/wp-content/uploads/2020/02/ECP_ST_SOLLVE.pdf
[vi] https://github.com/SOLLVE/llvm-project
[vii] https://legion.stanford.edu/pdfs/legate-preprint.pdf