Applying Graph Algorithms to Solve Key Science Problems of Importance to the Nation
By Scott Gibson

Clockwise from top left: Mahantesh Halappanavar, Pacific Northwest National Laboratory (PNNL); Aydin Buluc, Lawrence Berkeley National Laboratory; Marco Minutoli, PNNL; and Alex Pothen, Purdue University; not pictured: Erik Boman, Sandia National Laboratories
In this podcast, we explore the efforts of the Department of Energy’s Exascale Computing Project (ECP)—from the development challenges and achievements to the ultimate expected impact of exascale computing on society.
The ExaGraph project is what’s known as an ECP co-design center. ExaGraph is working on the efficient implementation of graph algorithms of key importance to the nation. We’ll get into what those things mean and shine a light on just how widespread and significant graph algorithms are in our lives. Specifically, subject-matter experts will give us a high-level view of what ExaGraph is all about.
Joining us are ExaGraph principal investigators Mahantesh Halappanavar of Pacific Northwest National Laboratory (PNNL), Alex Pothen of Purdue University, Aydin Buluc of Lawrence Berkeley National Laboratory, Erik Boman of Sandia National Laboratories, and Marco Minutoli of PNNL.
Our topics: the mission and activities of ExaGraph, a brief primer on graphs, what graph analytics can reveal, ExaGraph’s collaborations with ECP applications, and more.
Interview Transcript
Gibson: Mahantesh, what is ExaGraph and its overall mission, why did ECP fund it, and where does it fit into the big picture of ECP?
Halappanavar: Thank you, Scott. The overall mission of ExaGraph is to identify the key combinatorial optimization problems—and we’ll come to it in a little bit—that emerge in exascale application projects that are funded in ECP and develop scalable implementations that target both the current pre-exascale systems as well as the future exascale computing platforms.
ExaGraph is one of the six co-design centers under the Application Development thrust in the Exascale Computing Project. Co-design projects target cross-cutting algorithmic methods that capture the most common patterns of computation and communication, and these are collectively called motifs. Our motif was targeting graphs, graph algorithms.
Gibson: Please tell us about graphs. That is, what are they and how are they used. A little something about both the history and modern use will be helpful, I believe.
Halappanavar: Yes, definitely. Roughly 285 years ago, one of the greatest mathematicians of history, Leonhard Euler, posed an interesting puzzle known as the Seven Bridges of Königsberg. Two large islands in the city of Königsberg, Prussia, were connected to each other and the mainland by seven bridges across the Pregel River. The puzzle was to check if it was possible to follow a path that crossed each bridge exactly once and returned back to the starting point. And the solution to this problem is usually considered to be one of the first theorems in graph theory.
So, what are graphs? They are simple mathematical abstractions consisting of a pair of sets. The first entry in this is a set of vertices that represent unique entities, and the second is the set of edges, where each edge represents a pairwise relationship on vertices. For example, if you have a social network, people form the set of vertices, and an edge can represent some kind of relationship that could be something like a friendship between two people in the network.
Gibson: The branch of data science known as graph analytics is used to gain insights into many areas of society. What sort of knowledge can graph analytics provide, and how do the methods and techniques that the ExaGraph team is developing harness the capabilities of that form of analysis?
Halappanavar: You know, these days one does not have to be a computer scientist to notice how algorithms are all around us. They determine almost all aspects of our daily lives—the colleges we end up going to, the routes we end up taking based on the suggestions by our GPS systems, the schedules that determine our trains and our airplanes and our buses. They’re all based on algorithms. And even suggestions on who we should be friends with or what kinds of products we want to buy, so, again, they’re also recommended by projects. Many of them can be modeled as graph algorithms and solved efficiently using graph algorithms.
One such example, is the algorithm called the Match. This is executed by the National Resident Matching Program, or the NRMP. The Match determines the medical residency programs [in hospitals] that tens of thousands of medical school students and graduates from across the world get to be placed in each year. The Match is just one example of the algorithms that are being targeted for scalable development in ExaGraph.
Gibson: Thanks, Mahantesh. Now let’s turn to Alex for responses to the next few questions. First, Alex, what is combinatorial scientific computing, and how is the work of ExaGraph related to it?
Pothen: First of all, thank you, Scott, for this opportunity and thank you, Mahantesh, for organizing this. Much of scientific computing happens in the realm of what is called continuous mathematics—things like differential equations, matrix computations, statistical models, etcetera. But in today’s computing world, much of it also happens in the realm of discrete mathematics, or of objects like networks or combinatorial graphs that Mahantesh talked about. And of objects that can be counted like 1, 2, 3, like integers rather than 1.23 and so on, not a real number.
Researchers in combinatorial scientific computing identify and formulate problems in combinatorial discrete mathematics that arise in scientific computing and then they design algorithms to solve these problems, implement them on high-performance computers, and deploy them to solve problems in science and engineering.
A pioneering CSC, combinatorial scientific computing, research institute was funded by the [Department of Energy] Office of Science out of the SciDAC program—Scientific Discovery through Advanced Scientific Computing—and was called CSCAPES [Combinatorial Scientific Computing and Petascale Simulations] from 2006 to 2011 for developing combinatorial algorithms to enable the solution of scientific computing problems on petascale computers. ExaGraph is a very natural offshoot of the research efforts in this community, combinatorial scientific computing, to solve problems that have to do with graph modeling and combinatorial modeling on exascale computers.
Gibson: In some cases, the ExaGraph team has created parallel versions of algorithms they wanted to explore that actually didn’t exist before. Will you share with us some about that initiative?
Pothen: Many of the combinatorial problems that we work on require fairly sophisticated algorithms to solve them and don’t offer much concurrency, or parallelism. What we have to do is develop approximation methods that don’t solve the problems exactly but give us an approximate solution that is within some constant factor, say, a factor of 2, of the best possible solution. But these approximation algorithms have the advantage that the amount of work involved in solving this problem on a parallel computer grows only linearly in the size of the problem. If you have, say, a graph with a certain number of edges or a matrix with a certain number of nonzero elements and so on, the work is only nonlinear in the size of that metric. Also, many of these approximation algorithms can be designed to have much more parallelism than the exact algorithm, so we can actually solve them on very large computers.
The ExaGraph team has explored this paradigm of using approximation algorithms to develop solutions that can be computed in parallel effectively. And an instance of this is the matching problem that Mahantesh alluded to. The matching problem comes up in the context of developing a significant subgraph, for example, of a much larger graph or a much larger data set, and for downstream scientific computing, or inference tests.
We have developed algorithms by this approximation technique that scales to 16,000 processors of a distributed memory computer to solve problems such as the data privacy problem and so on. And that’s just one instance of developing these parallel algorithms.
Gibson: Many of our listeners may already be aware that ECP has a project called NWChemEx. It’s developing high-performance computational “models demonstrating that biomass can be a viable, sustainable feedstock” for the production of biofuels and other bioproducts. And I know that ExaGraph has supported NWChemEx. Please share with us about those efforts.
Pothen: Sure, Scott. The NWChemEx computation is using quantum chemistry to model large molecules like proteins and so on and understanding the energy levels so they can understand their chemical and biological properties. In their computation, they need to compute a matrix called a Fock matrix to do what is called density functional theory in quantum chemistry.
If you can think of the number of atoms that you use, and something related to that, and call that a variable n. Then the amount of computation involved in computing the Fock matrix grows like the fourth power—that is, n4. That is n × n × n × n. It grows that quickly, and so, this is one of the most demanding steps in this computation. NWChemEx uses a supercomputer at Oak Ridge National Laboratory called Summit, one of the leading computers in the world today in terms of supercomputing ability. They use this computer to do this step, and they need to minimize the computation time that the processors take by assigning work in a balanced way to the processors and also making sure that the processors send roughly the same amount of messages using a messaging passing library called MPI.
We formulated this problem as a matching problem, but using a nonlinear objective function called a submodular function rather than a linear function, which is what people commonly do. Submodular functions describe the property of what are called diminishing returns. That is, suppose we have two bags of tasks to do, and think of two subsets of this bag of tasks [A and B]. Let’s say we have a small subset [A] and a big subset [B], and the big subset contains the small subset [A ⊂ B]. Suppose I take a new task and I assign it to the small subset [A]. How much will it improve the objective function? If the improvement is better than when you add it to the bigger subset [B], we have a submodular function. It’s a property of diminishing returns—the gain that you get by adding an element to a large set is smaller than what you’d get by adding it to a smaller set to begin with.
By using this formulation, we were able to balance a workload on each processor and also minimize the number of messages sent, and we speeded up the computation by a factor of 4 over the current default load-balancing scheme that is used in NWChemEx. We also scaled the computations to 8,000 processors instead of the 3,000 processors that they were able to scale to, using, again, the default load-balancing scheme. Now a computation that use to take more than 30 minutes, can be done in about 8 minutes.
Our paper on this topic will appear in the Proceedings of the first SIAM Conference in Applied Computational Discrete Algorithms, which will be held this July. This summer we are continuing to work with the NWChemEx team to make sure that we can continue to load balance their computations on larger numbers of processors and also provide more accurate algorithms so that load balancing is more effective.
Gibson: Now over to Aydin . . . will you share with us about the ECP applications that ExaGraph is working with?
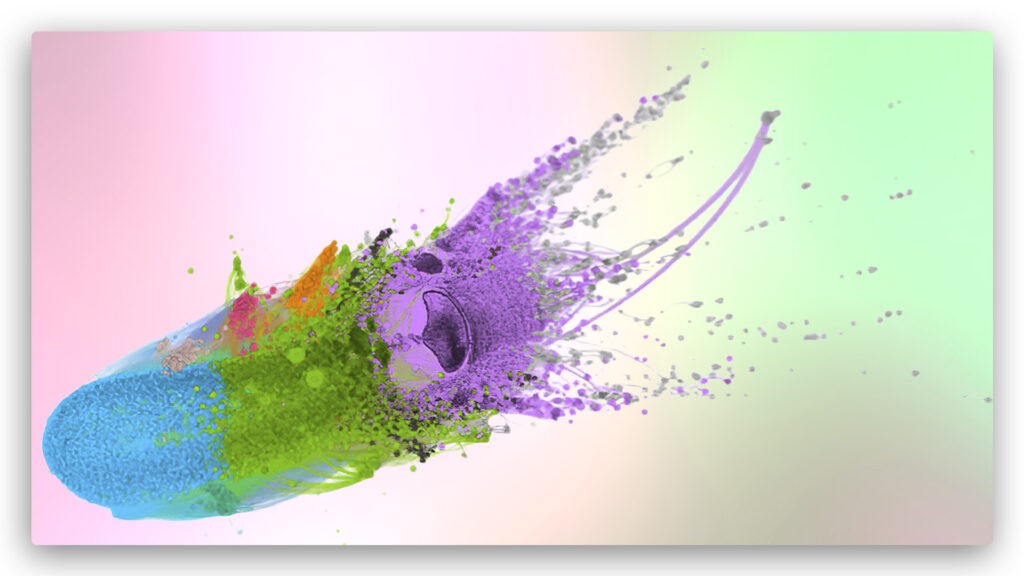

Proteins are the workhorses of life. Protein functions can be determined from protein sequences (structure). High-throughput sequencing of genomes and metagenomes has dramatically expanded the sequence data that is now available. However, function for a large fraction of proteins is currently unknown, and therefore, computational annotation of metagenomes has become a critical task. Construction of protein similarity graphs (networks) enables efficient prediction of functions. The figure illustrates the rendering of a protein similarity network constructed from the PATRIC database with 170K vertices (proteins) and 1.7 million edges, where colors indicate clusters (or communities) of proteins. Only the largest component is show in the figure. Credit: the ExaGraph co-design center, Exascale Computing Project
Buluc: Yes, my pleasure. Alex already gave a lengthy response to ExaGraph’s relationship to an ECP application, NWChem; therefore, I’m not going to repeat it. Other applications we support . . . a primary one that I am involved with is ExaBiome, which works on various biology problems and exascale. I will mention a little bit of that later in the podcast. Otherwise, ExaGraph works with several national security applications and that usually stems from the fact that for many engineering and science problems, the fundamental operation is solving a system of equations, and often these are linear equations or there’s an iterative algorithm that eventually solves a linear equation.
And these are the same things that you do in linear algebra in freshman or sophomore [year of high school]. You just write X + Y + Z = some value and then say X – Y + 2Z = some other value. Think of this multiplied with millions. These are the systems that engineers and scientists solve daily in various applications, including some of national security applications in ECP.
As the systems get more specialized—meaning when they get sparse—that means that several of the coefficients are zero. For example, the coefficient of X is not 2, but it’s 0. The coefficient of Z is not 3, but it’s 0. Those systems become sparse, and it, of course, makes a lot of sense to take advantage of that sparsity. And some of the methods that ExaGraph has developed in [graph] matching directly can be applied to these problems. These are not just national security applications; some of them are science applications.
There’s also a specialized solvers methodology called a multigrid, and there you’re successively trying to create a system that looks like the original system, but it’s smaller and captures the major characteristics. You do this in a recursive manner, meaning you get a smaller system and from that smaller system you get an even smaller system, and so on until it’s small enough that you can solve it and project that solution back. And there’s another important problem ExaGraph is working on, graph coloring, that comes into play where we’re figuring out what parts of the equations can be coarsened together. Or what is the right way of coarsening this system to get a smaller system?
The final kind of application that comes up often is also another security application where we need to compute derivatives. Derivatives of a function—as anybody who has studied a little bit about calculus knows—are needed in an enormous number of science and engineering applications. More recently it’s been also used a lot, thanks to the popularization of machine learning—like deep learning, where derivatives are very important. A Jacobian is just a matrix form of the derivatives of a function, and computing those involves a particular system. Derivatives can also be sparse, just like the equations we just talked about can be sparse, and at that point, the same coloring problems are applied to these applications. And I can, of course, give many, many more examples, but I think this is already a long enough answer.
Gibson: OK. You discussed ExaBiome, let’s turn our attention to that project. They’re identifying groups of similar proteins, and ExaGraph has teamed up with ExaBiome. Will you tell us about that collaboration?
Buluc: Yes, happily. As you might know, I’m at Berkeley, so it was a little natural, that collaboration that I brought into ExaGraph, and I’m also part of the ExaBiome team. ExaBiome works on various different problems in computational biology, and one of them is identifying groups of similar proteins, as you said. Similarity between proteins signifies something known as homology, and all that really means is those proteins evolved from a common recent ancestor. The practical implication of that is that those proteins that evolved from a common ancestor usually perform the same function, and the simple corollary of that is if I know protein A, already in my database—I worked on it, I studied it, and this takes a lot of time—and I see another protein that I just sequenced from an environmental sample, and I don’t know what it does; it looks like it’s similar to the thing I know in my database. I can say, OK, this probably does the same function. Let me do a quick check, and, yes, and move on. That’s, of course, a lifesaver.
However, the more important corollary of that is, what if I get a group of similar proteins and none of its members include anything that I know of? And that happens, especially when we get samples from remote corners of the earth, such as a soil sample or a sample from a mud land, a sample from an ocean. These can tell me new families of proteins that contain no known members to the scientific community. These are novel proteins, and their functionality can be very useful for a lot of pharmaceutical needs or others. For example, if I find a new protein that breaks down certain crops faster than any of the enzymes that I know, then that protein can be used to produce biofuels in a much more efficient way. That’s really the whole goal of finding similar proteins.
ExaGraph helps on this process by working on the whole pipeline, which involves first building a graph—and we already talked about what a graph is—where the edges in the graph are just similarities between the proteins and the vertices are the proteins themselves. From then on, it builds a clustering of those using a method known as a flow, or a random walk, and then those clusters are deemed to be the protein families.
Now, one of the reasons that this work is done in collaboration with ExaGraph is because both the graph clustering that is downstream as well as constructing a similarity graph from a dataset have applications beyond biology. It makes sense not to just leave the software in a biology project repository but also expose it to a broader set of both ECP and beyond-ECP science applications.
Just to give an example, after we publish the HipMCL [the graph clustering work] paper, half of the citations that we get for that work is, of course, by the biologists that use if, but the other half is completely outside the biology domain. We get citations from people that work on routing on the internet where they have clustering problems, and network security guys that cite and use our software. The applications are broader than just biology and it made sense to bring it to a co-design center as opposed to leaving it in an application project.
Gibson: Wow, the broad applicability of ExaGraph’s published knowledge is interesting. I believe Erik has some insights for us. Erik, what is ExaGraph’s main way of delivering software?
Boman: Yeah, Scott. That’s a good question. First of all, I should say all our software is open source and made available to the HPC [high-performance computing] community. In fact, we put it on websites like Github so anybody, even beyond ECP, can download it. Now, there is not a single software package called ExaGraph. ExaGraph is a project, and we have multiple ways to deliver the software because there are several labs and universities involved, and we all work with different applications. It actually seemed better to deliver the software in such a way that it’s easier for the applications to use the software.
There’s not a simple answer, so I will focus on what we do at Sandia—I’m at Sandia National Labs. We have a large software framework called Trilinos to do scientific computing. Trilinos has many packages. One of them is called Zoltan2. It is a package for graph algorithms and combinatorial scientific computing. The main focus is on graph partitioning, which I can explain later. Our software that we have developed for graph partitioning is put inside Zoltan2, which means it’s part of Trilinos. Trilinos is part of ECP, so it’s also part of xSDK, the Extreme-scale Software Development Kit, which is used by many of the ECP projects. Therefore, our software is easily available to ECP applications.
Gibson: Let’s look at another term that’s germane to ExaGraph—graph partitioning. What is it, why is it important, and what advances in that area has ExaGraph made?
Boman: Graphs are really the mathematical abstraction for a lot of computing tasks, and graph partitioning is the problem of how to break up a large graph into many smaller graphs. Typically, when we do parallel computing the whole problem cannot fit on a single processor, so we have to partition it and divide it into smaller pieces so each smaller piece can be processed by a single computer or compute node. It’s really an optimization problem where the constraint is a balance constraint that you want to split up your graph into approximately equal pieces because this corresponds to the workload and then assign it to processors, but you also have to minimize the interaction between the processors because that will generate communication, which is expensive in parallel computing.
From a graph perspective . . . and remember every graph has vertices and edges, so you want roughly the same number of vertices on each processor, and you want to minimize the number of edges that go between processors. This is the graph partitioning problem. This problem has several variations, but the balanced partitioning is the one that is most relevant to scientific computing and parallel computing in particular. So that’s what we have focused on. And even that problem has been studied for several decades.
There was a lot of work done back in the 1990s, and the algorithms developed there are still used today. One of our contributions is really focused on the ECP computer architectures. One difference is that we now have computers with the GPUs and other accelerators in the parallel context. We have multi-GPU computers and the current graph partitioners. So there are already algorithms and software for graph partitioning, but they were developed years ago for CPUs, the old-style computers, while now we need to focus on GPUs, the graphics processing units.
Our main contribution is a new partitioner—we call it Sphynx—and what it does is that it implements a graph partitioning algorithm based on what’s called spectral partitioning, which uses linear algebra and was first developed actually by Alex Pothen and his group. We have implemented this in the context of Trilinos to use GPUs and CPUs, so the same code can run on both GPU-based machines and CPU-based machines, which previous software could not do. This should enable applications to run efficiently on the new exascale computers that heavily rely on the GPU, the graphics processors.
Gibson: Excellent. Thank you! Marco is going to help us with the next question. ExaGraph has achieved much in the pursuit of a type of problem known as influence maximization. What’s involved in those activities?
Minutoli: Let’s start with understanding first what is influence maximization. The problem was posed in the context of viral marketing in the beginning, and the idea in viral marketing is to optimize so we could distribute free samples and then leverage the word of mouth and recommendations between people and actually spread the adoption of some product. In this context when the original problem was posed, the question was, how do we choose a small set individuals from a population or a social network so that we could give them free samples and optimize their word of mouth process of diffusion?
The task is actually asking to select from a broader population of a small set of individuals that we optimize in the diffusion process, and that means some process that we either carry information within the network or can model a lot things like, for example, how diseases are spreading over a network or other preferences over some problem, as I said before. Or, for example, we can choose ways of deploying sensors in a harsh environment and things like that.
As a computer scientist, the way I see it our role is to provide tools that will enable other scientific discoveries. At the time when we started with our work on influence maximization, even though there were already breakthroughs in the field that allowed us to formulate the problem in an efficient optimization framework like the one that Alex has described, the submodular optimization framework, the tools that were available were not really scalable and still limited in the size of the graph and problem instances that we were able to solve.
What we have done across the years is to actually develop parallel algorithms that can leverage supercomputers at the US Department of Energy essentially to provide more efficiently a solution to this problem. We designed the first parallel algorithm for this problem, leveraging some work done by other scientists, and with that implementation we moved from, essentially, hours of computation for a specific instance of this problem to roughly 50 seconds of computation or an hour over a thousand processors on a big, distributed machine. That’s quite a big improvement.
Our later work was actually on reducing the size of the system need to efficiently solve these problems. And we started looking at GPU optimization on our next generation of exascale computer with the GPUs, so we designed an algorithm for those and we reduced the number of nodes of the class that we needed to solve the same instances that at the beginning were taking roughly 8 hours with only 64 nodes on Summit, which is the supercomputer that was mentioned before.
The last contribution we did actually is to tie a connection between the problem of maximizing the influence over a social network and the problem of deploying, for example, vaccines using the framework for network epidemiology, which is a framework based on the network science, where we are modeling interactions between people and trying to give them immunity through vaccination. And we designed a pre-emptive strategy so that when you know that something is coming up, you vaccinate in the network. That is giving us a best-case scenario of what we could actually achieve. To do that, we leveraged the concept of maximizing the number of people that are not getting the disease, which is very close to the idea of influence maximization, and we use the algorithms that we have designed in the past. These are the problems.
Gibson: Thanks, Marco. As we wrap up, let’s turn our attention to what the ExaGraph team is focused on now and what your plans are for the near future. Mahantesh?
Halappanavar: Current work can be categorized along three components: algorithmic development, scalable implementations, and applications—exascale applications in particular. Future work will continue along these three components, and, in particular, we’ll be focused on optimization and porting to the impending exascale computing platforms. Since the new platforms have several different hardware components from different hardware vendors—and therefore, diverse computing environments—porting to these new platforms is nontrivial, and we expect, as such, to spend quite a bit of effort trying to make the code optimal in performance as best as we can get.
Another aspect that we want really to focus on is to work very closely with exascale applications and enable their applications and algorithms that are developed in ExaGraph. And what we have presented here through Aydin, Alex, Erik, and Marco was just a small portion of the entire work that’s being done under ExaGraph. We do have a large number of participants. I sincerely thank every one of the participants in ExaGraph and conclude by saying that we want to leave a lasting legacy of algorithms, implementations, and graph-enabled applications through ExaGraph. Thank you.
Buluc: It’s all an exciting time to be working with everybody in ExaGraph and lots of exciting applications across the ECP portfolio.
Pothen: I’ll add that I am one of the few non-[national]-lab participants in ExaGraph, and I’m very grateful for that opportunity. And I hope that future opportunities for doing research and for having such a large project will definitely include other university participants as well, because I think that we could add some value to the work that is being done at the labs.
Boman: I agree with what Alex just said. In fact, we do have collaborations with other university partners as well as. I only focused on one partner in the work at Sandia. We are also working on a multilevel algorithm. This is what Aydin talked about before. We have a large problem making smaller problems that are similar, and that’s a quite promising approach especially for graph partitioning. And we have an active collaboration with Penn State, Professor Kamesh Madduri, and I definitely think university partners have a lot to contribute in the context of ECP.
Minutoli: As a young researcher working in a national laboratory, ExaGraph has been a great opportunity for me to grow and actually meet other scientists from other laboratories, grow my network, and start collaborations that I think will be essential to get more exciting work in coming years. And, yeah, I honestly hope that in the future we’ll see more of this kind of projects and collaboration between lab, industry, and universities that will make things more exciting.
Gibson: Thank you all for being here.
Halappanavar: Thank you very much, Scott.
Related Links
- ExaGraph project details on ExascaleProject.org
- ExaGraph co-design center website